Instrumentation
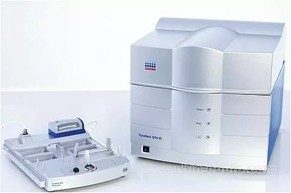
PyroMark Q96 ID
The PyroMark Q96 uses pyrosequencing technology combined with a 96-well format to provide quantitative analysis of genetic and epigenetic DNA modifications as well as mutation gene expression analysis. It has an automatic base-calling function and dedicated software to aid methylation assay design and analysis.

Illumina MiSeq System
The MiSeq system is used for in-house DNA methylation analysis and QC for ChIP libraries. It is capable of automated paired-end reads and creates up to 15 Gb of output with 25 million sequencing reads and 2 × 300 bp read lengths per run.

Oxford Nanopore GridION
The GridION Mk1 can directly detect modified bases in nucleic acids without intermediate processing. This feature, combined with its ability to generate several kilobases of nucleic acid sequence per sequencing pore, makes it useful in many epigenomic analyses.

Covaris ME220 Focused-Ultrasonicator
The Covaris ME220 Focused-ultrasonicator is a multi-sample, multi-application benchtop sonicator formatted for 1 to 8 sample batch-processing. It is equipped with an integrated chiller and has automated water management. It helps ensure accurate, reproducible and robust sample preparation.

Bioruptor Sonicators
A key step for ChIP and next-generation sequencing experiments is precisely controlling shearing of chromatin, DNA and RNA into appropriately sized fragments. The core possesses a Diagenode Bioruptor Twin and a Bioruptor Pico sonicators, which are all-in-one, water-based, shearing device with advanced temperature control. These devices use highly-controlled ultrasonic energy with ACT (Adaptive Cavitation Technology), a non-contact, parallel-processing and isothermal technology, to achieve optimal fragment sizes between 150bp-2kb.

Agilent 2100 Bioanalyzer
The Agilient 2100 Bioanalyzer performs electrophoresis in microchannels to provide sizing, quantitation and purity assessments for DNA and RNA samples.

Agilent 4200 TapeStation
The Agilent 4200 TapeStation is an automated system providing a rapid, reliable gel electrophoresis platform to assess overall sample quality (e.g. size, quantity and quality) for 1-96 samples prior to next-generation sequencing (NGS), microarray (aCGH) or quantitative PCR (qPCR) applications.

QuantStudio™ 6 Pro Real-Time PCR System
Give Now
Research Areas
Find out about the four types of research taking place at UT MD Anderson.
