Current Research
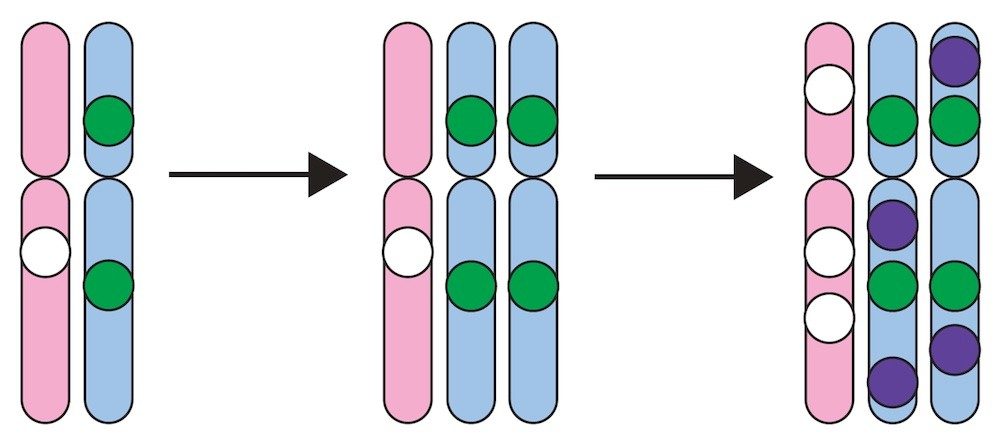
Molecular archeology of cancer
The cancer genome contains an archeological record of its past. The
Cancer Genomics and Evolution Laboratory has pioneered methods to
reconstruct a cancer's life history from massively parallel sequencing
data and has used these approaches to obtain detailed timelines of
tumor evolution across many cancer types.
While we now
have detailed insight into the drivers, mutational processes and
evolutionary timelines of primary tumor development, in comparison,
our understanding of the metastatic process is more limited.
Nevertheless, metastases cause 90% of cancer-related deaths. The
Cancer Genomics and Evolution Laboratory is now using existing
molecular archeology of cancer approaches, as well as developing new
ones, to reconstruct the evolutionary history of cancer metastases.
Through this, we aim to get new insights into metastatic
dissemination, and identify new leads for early diagnosis and
treatment stratification.
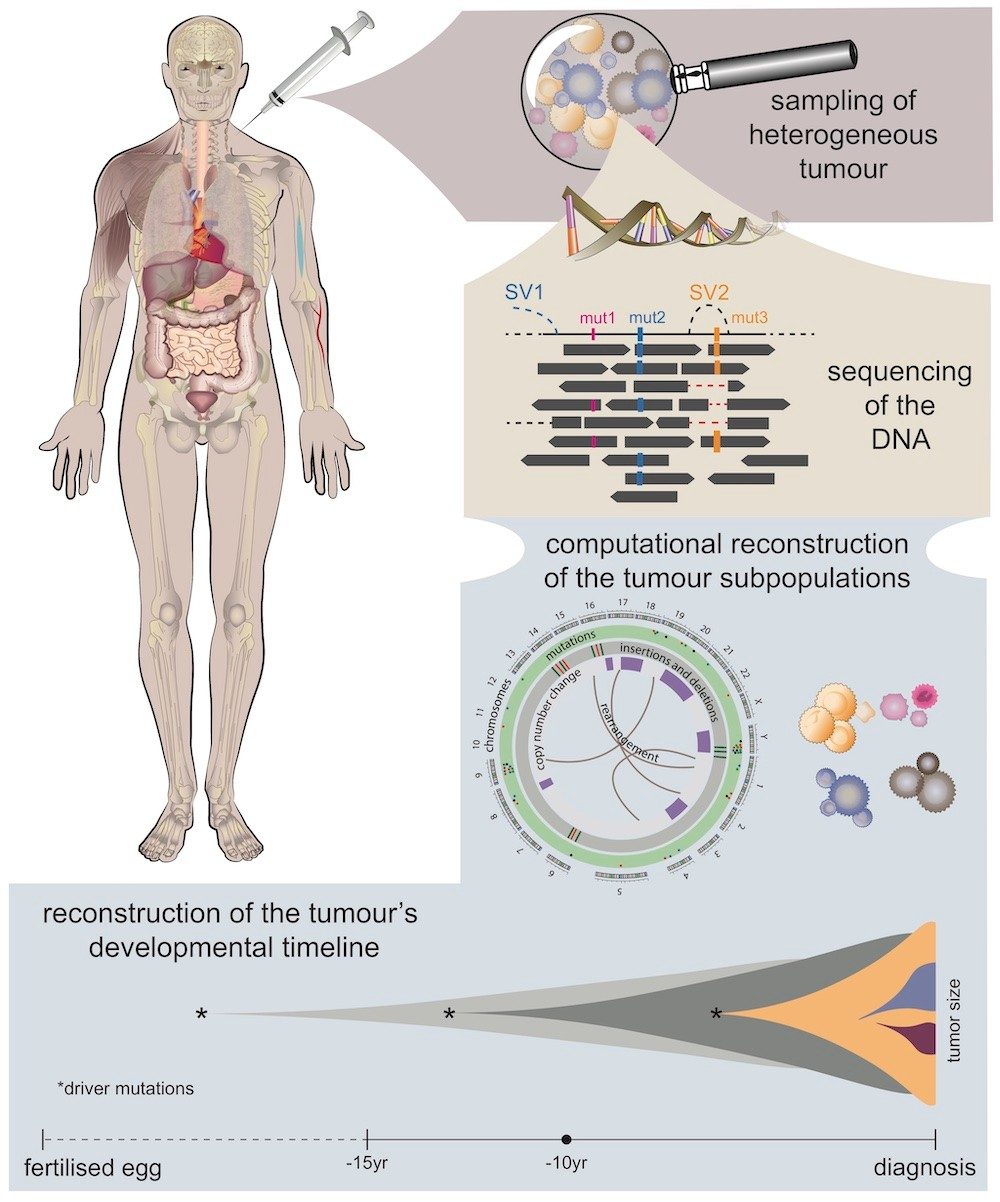
Intra-tumor heterogeneity through the lens of bulk and single-cell sequencing
Tumors are heterogeneous mixes of subclones, and this intra-tumor
heterogeneity fuels tumor progression and treatment resistance. The
subclonal architecture of cancer can be profiled in multiple ways,
including bulk tumor sequencing and single-cell sequencing.
To study intra-tumor heterogeneity in minute detail, the Cancer
Genomics and Evolution Laboratory is using bulk whole genome
sequencing, combined with novel single-cell and spatial sequencing
approaches. We develop computational and statistical methods to
integrate data across these different technologies and thereby
reconstruct the subclonal architecture of tumors in minute detail. We
apply these approaches across primary tumors, cancer recurrences,
disseminated tumor cells, and cancer metastases, aiming to get insight
into tumor evolution, response to treatment, and metastatic
dissemination.
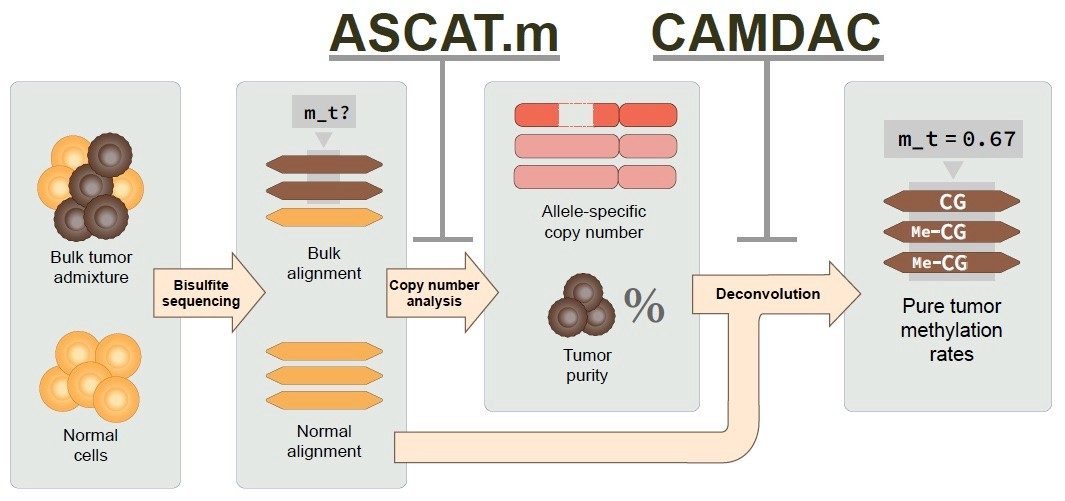
Molecular archeology of the cancer methylome
As epigenomic changes are inherited over cell divisions, they can
also be used to track a tumor’s evolutionary history. We have recently
developed a tumor-normal deconvolution approach for methylation data
(bisulfite sequencing), leveraging tumor purity, allele-specific copy
number, and an adjacent normal sample as a proxy for admixed non-tumor
cells. Using this method, we are now able to study differential
methylation and intra-tumor heterogeneity at the methylation level in
cancers.
The Cancer Genomics and Evolution Laboratory is
developing molecular archeology of cancer approaches to time
methylation changes in tumor evolution, and to use methylation changes
to time other (epi)genomic events. These methods hold the potential to
both significantly increase the accuracy with which we can time
events, and considerably increase the breadth of events we can time.
Through these approaches, we aim to shed light on the evolutionary
history of both the cancer genome and epigenome, giving us insight
into how cancers develop, evolve and metastasize.
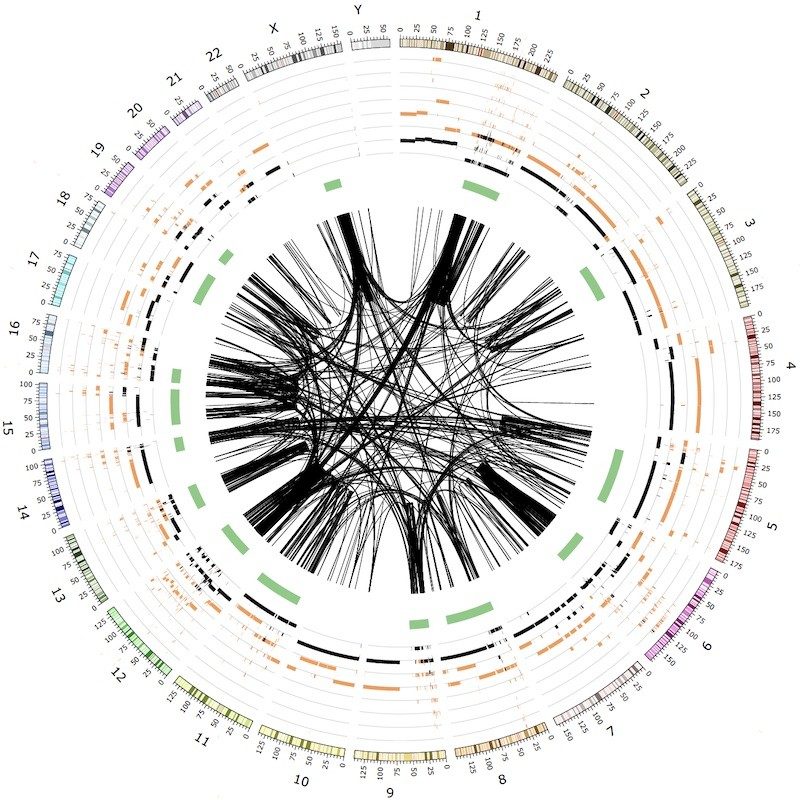
Cancer genes and mutational processes
The mutational processes and cancer genes driving tumor
development can vary across cancer types. Massively parallel
sequencing studies have identified many frequently mutated cancer
genes, and many underlying endogenous and exogenous processes
causing mutations. Nevertheless, there are still significant gaps,
such as rare cancer genes mutated in highly specific cancer
(sub)types, and in mutational processes driving complex structural
variants and copy number changes.
The Cancer Genomics
and Evolution Laboratory is leveraging multiple genomic profiling
approaches to identify novel rare cancer genes, and to identify,
characterize and time mutational processes causing copy number
changes and complex structural variants in cancer evolution. We have
a particular interest in sarcomas, which often feature complex
structural variants, and which, as a collection of many individually
rare cancers, may harbor yet undiscovered recurrent structural
variant or point mutation drivers.
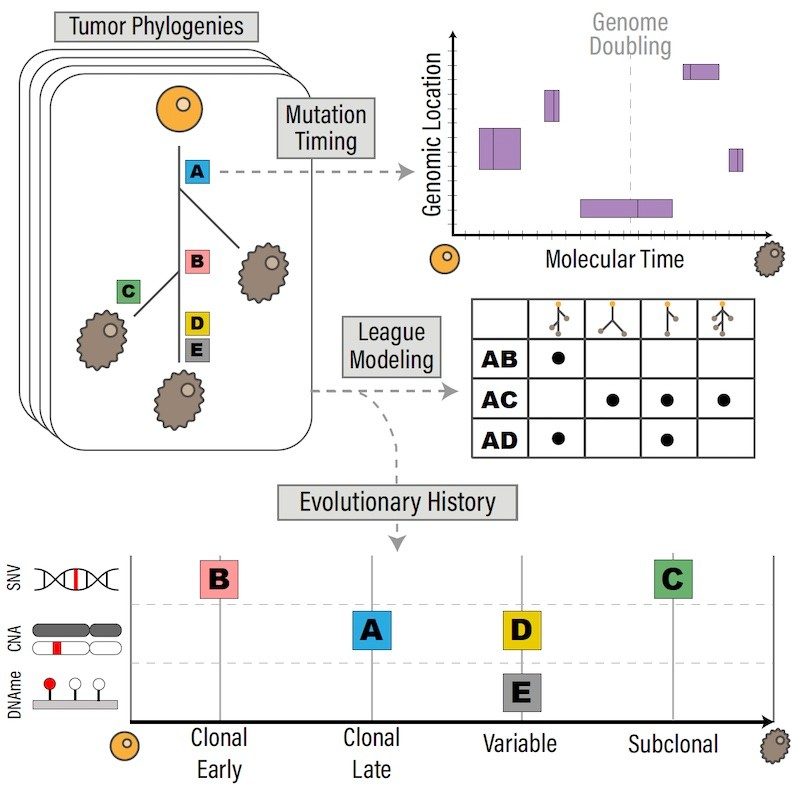
The evolutionary history of metastasizing gastrointestinal cancers
Metastases of colorectal and pancreatic cancers represent the second and third most common cause of cancer deaths in the US. The Cancer Genomics and Evolution Laboratory leverages whole-genome sequencing, whole-genome methylation profiling, the institutional post-mortem tissue collection program, and its molecular archeology of cancer approaches to understand the evolutionary history of colorectal and pancreatic cancers. Combining molecular archeology of cancer approaches at the genome and methylome level, we will construct detailed evolutionary trajectories, including both genomic and epigenomic events. Through this, we aim to get new insights into the metastatic process in these cancer types, allowing us to identify when key genomic and epigenomic changes occur in tumor evolution, and what their roles are in driving metastasis.
Give Now
Research Areas
Find out about the four types of research taking place at UT MD Anderson.
