Research
Epigenetic gene regulation is important for both normal development and for creating disease states. In cancers, aberrant promoter CpG island hypermethylation correlates highly with gene inactivation and can account for a loss of gene expression despite the lack of genetic mutation. Dr. Ting is interested in dissecting the mechanisms of epigenetic gene silencing and understanding the functional relevance of DNA methylation in diseases. Dr. Ting’s lab has four major focus areas: 1) pioneering technical and computational tools for genome-wide DNA methylation assessment; 2) defining novel and clinically relevant functions for DNA methylation; 3) delineating the mechanics of DNA methylation in cancer; and 4) GenitoUrinary Development Molecular Anatomy Project (GUDMAP) Atlas development.

- Pioneering technical and computational tools for genome-wide DNA methylation assessment
Dr. Ting developed methyl CpG binding domain (MBD)-isolated Genome Sequencing (MiGS), a cost-effective technique to survey whole genome DNA methylation patterns. The Ting lab also develops computational tools that facilitate sequencing data analysis and interpretation.
- Defining novel and clinically relevant functions for DNA methylation
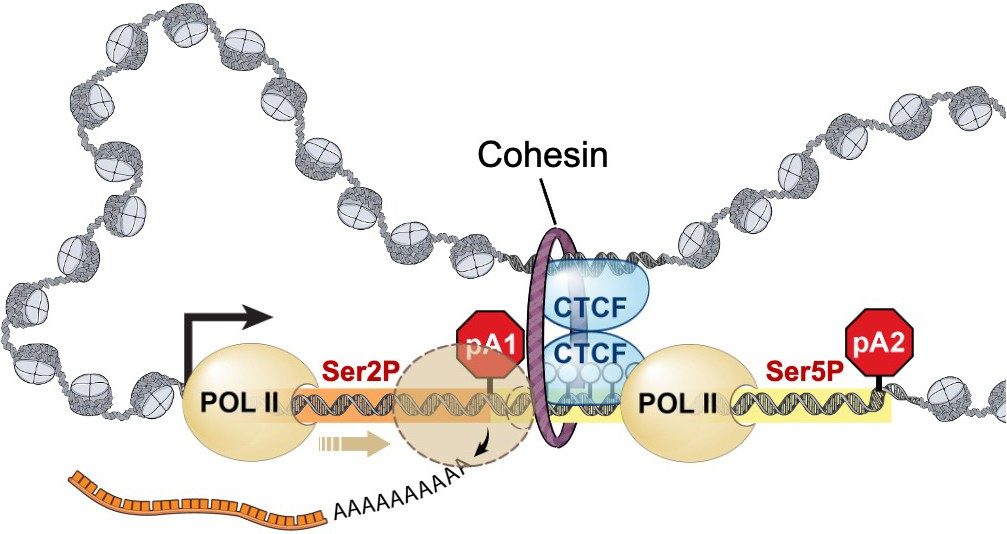
The Ting lab has defined genome-wide abnormal DNA methylation patterns in colon, bladder, and prostate cancers. Knowing where these disruptions occur enabled her group to generate and test hypotheses regarding the functional consequences of these changes. They are now investigating both gene promoter and non-promoter regions with the goal of defining context-specific functions of DNA methylation such as the lab's recent discovery, published in Molecular Cell, showing that DNA methylation regulates alternative polyadenylation via CTCF and the cohesin complex.

- Delineating the mechanics of DNA methylation in cancer
Wide-spread disruptions to DNA methylation patterns contribute to tumorigenesis and progression, but the regulatory mechanisms that establish, maintain, and modify these patterns remain unclear. Researchers in Dr. Ting’s lab explore the roles of epigenetic enzymes, small non-coding RNAs, and microbiome-host interactions in the initiation and maintenance of abnormal DNA methylation patterns in cancer. For example, Dr. Ting is investigating whether enzymes such as methyltransferases (MTases) or restriction enzymes (REs) expressed by bacteria present in the microbiome and/or tumor microenvironment can directly modify/cut host DNA to drive cancer.
- GenitoUrinary Development Molecular Anatomy Project (GUDMAP) Atlas development
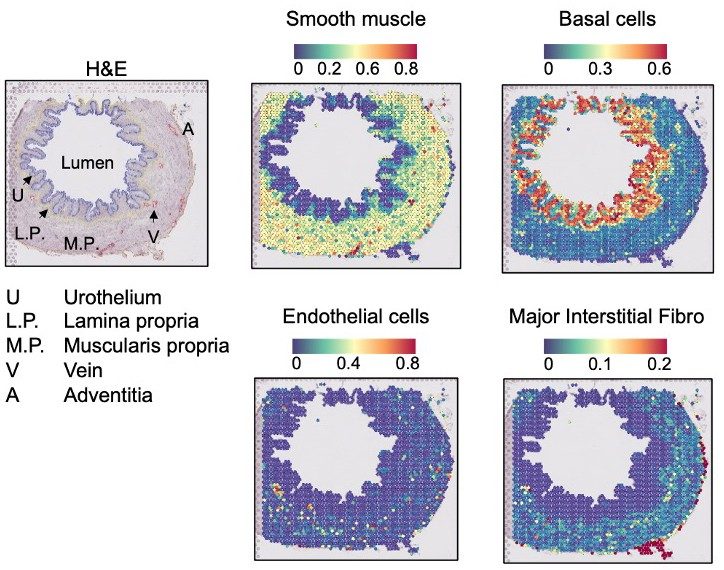
The Ting lab, together with Byron Lee, M.D., Ph.D., also at MD Anderson, and Oliver Wessely, Ph.D., at Cleveland Clinic, are generating a cellular and molecular map of the normal human ureter and bladder as part of the National Institute of Diabetes and Digestive and Kidney Diseases GUDMAP consortium. Leveraging state-of-the-art single cell and spatial genomic platforms, they are identifying the cell types that make up the human lower urinary tract and defining how the transcriptomes of these cells are modulated by signaling pathways and epigenetic mechanisms. Portions of this work were published in Developmental Cell (see the GUDMAP Atlas data we generated as part of this work) and in iScience (see the GUDMAP Atlas data stemming from this work).
Give Now
Research Areas
Find out about the four types of research taking place at UT MD Anderson.
