Current Research
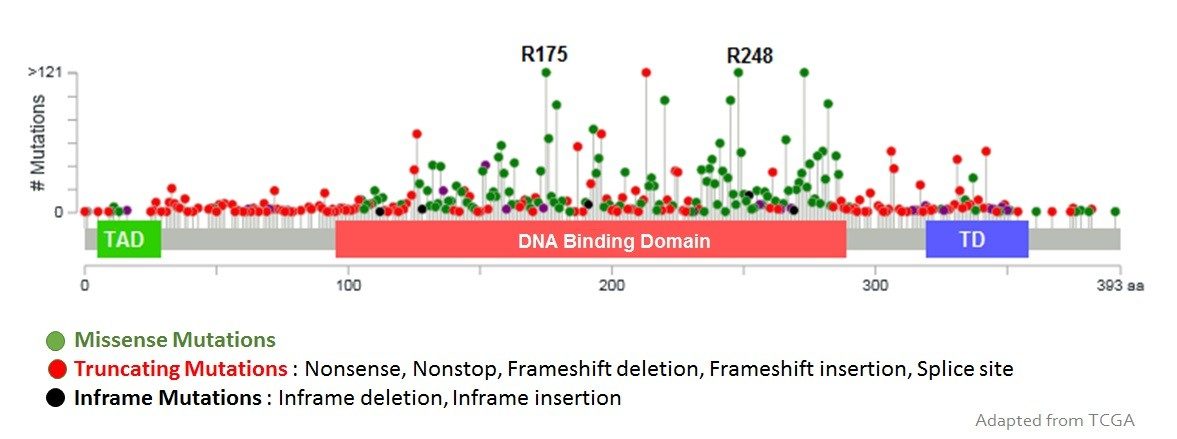
The research in my lab has been focused on understanding the functional mechanism of cancer associated genes by generating mouse models. We have a long-standing interest in p53 gene as the most mutated gene in cancer. Mutation of the p53 gene is a critical event in the elaboration of many tumors of diverse origin. The p53 protein is activated in response to DNA damage, serving as a checkpoint in the elimination or repair of cells with damaged DNA. Alterations in components of the p53 pathway, such as amplification of the Mdm2 gene, which encodes a p53 inhibitor, also contributes to tumorigenesis. The overall goal of my laboratory is to understand the signals that regulate the p53 pathway and the consequences of expressing wild-type or mutant p53.
Mouse models of Mdm2 and Mdm4 loss
We have made conditional loss-of-function alleles of Mdm2 and Mdm4 to access the role of p53 in different cell types to probe the importance of components of the p53 pathway in tumorigenesis. For example, loss of Mdm2 or Mdm4 (genes that encode p53 inhibitors) results in embryonic lethality that is completely rescued by concomitant loss of p53. These studies indicate the importance of regulating p53 activity.
Mouse models of p53 mutation
The first mice generated in our lab were those expressing a common p53 mutation identified in human cancers. Mice expressing the p53R172H mutant develop osteosarcomas and carcinomas that metastasized at very high frequency. This study also indicates that mutant p53 is inherently unstable in vivo suggesting that other genetic alterations in tumor cells stabilize p53.
We have generated mice with another interesting mutant that distinguishes the ability of p53 to induce apoptosis or cell cycle arrest. The p53R172P mutant cannot transactivate genes that induce apoptosis yet retains the ability to induce the p21 gene involved in cell cycle arrest. Importantly, this mutant shows delayed tumorigenesis suggesting that p53 activities other than apoptosis are also critical for tumor suppression. Tumors that arise in these mice are also genomically stable suggesting that the p53R172P mutant transactivates genes involved in maintaining a stable genome.
Tumor microenvironment
We have recently generated a conditional p53R172H and p53R245W mutant mice that allows us to better understand the mechanism by which mutant p53 exerts its cellular effects to modify the tumor microenvironment. In this genetic model of somatic mutant p53, in contrast to germline mutant p53 which harbor p53 mutation in whole body, the expression of mutant p53 is limited to tumor cells and the stromal cells are wildtype for p53. We are interested in understanding how the cross talk between mutant p53 cells and the stroma contribute to metastatic phenotype by modulating the tumor microenvironment. Such knowledge has the potential to identify new targets for therapy and provides new insight into potential immunotherapy interventions for tumors with mutant p53.
p53 restoration
We have also explored the clinical implications of restoring p53 activity in different contexts using a hypomorphic p53 allele that restores p53 gene function upon Cre-mediated recombination. Restoration of p53 has different effects in different contexts. Restoring p53 in tumors that lack p53 causes tumor regression. Restoring p53 in a mutant p53 background or in tumors with amplified Mdm2 promotes suppression of tumor growth but does not cause tumor regression. These data highlight the importance of context in p53 response.
Give Now
Research Areas
Find out about the four types of research taking place at UT MD Anderson.
