Research
The Junjie Chen Laboratory focuses on understanding molecular mechanisms underlying genomic instability and tumorigenesis. Maintenance of genome integrity following DNA damage requires the coordination of DNA repair with various cell-cycle checkpoints. The hope is that by understanding these DNA damage-responsive pathways, we will know how deregulation of them contributes to tumor initiation and/or progression and how to take advantage of this deregulation in cancer therapy. The laboratory has been studying DNA damage signaling and DNA repair pathways since 1999. We have identified and performed in-depth functional studies of many key cell-cycle checkpoint and DNA repair proteins in several DNA damage signaling and repair pathways.
With the increasing appreciation of intricacy of signaling pathways, it becomes evident that we need to go beyond our studies of individual proteins and pathways. We should achieve a comprehensive understanding of the network involved in DNA repair and determine how these proteins and pathways intersect, interact, communicate, coordinate, and collaborate for genome maintenance. Only by elucidating the complexity of DNA repair network will we be able to make meaningful and decisive contributions to cancer biology and treatment. With this in mind, we successfully carried out several genome-wide to medium- and small-scale network studies in various DNA repair and oncogenic signaling pathways. The goal is to combine our ability to conduct network analysis with our expertise in performing detailed mechanistic studies to establish physical and functional networks of DNA damage response, tumor suppressive and oncogenic pathways, which will facilitate the long-term goal of exploiting DNA repair and vulnerability in cancer to revolutionize treatment for cancer patients.
Research Strategy
Define DNA damage response pathways
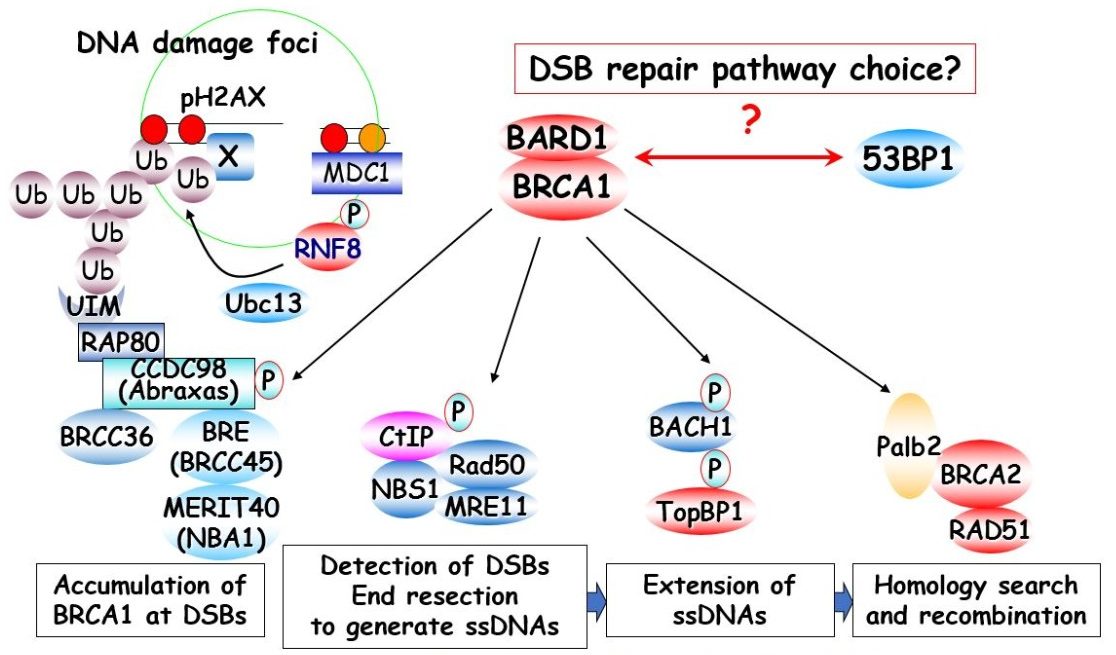
BRCA1 is the major familial breast cancer tumor suppressor. We showed early that BRCA1 associates and co-localizes with BRCA2 and the homologous recombination (HR) repair protein RAD51 and thus proposed that a defective HR repair pathway may be the underlying mechanism for the development of familial breast cancer. We have been studying the biological functions of BRCA1 and its precise roles in DNA damage checkpoint control and DNA repair for many years. BRCA1 has several conserved domains, including the BRCA1 C-terminal (BRCT) and N-terminal Really Interesting New Gene (RING) domains, the latter of which possesses E3 ubiquitin ligase activity. Our major contributions are a) finding that the BRCT domain is a phosphoprotein-binding domain and further demonstrating how BRCA1 associates and functions with several of its binding partners via this phosphorylation-dependent interaction, b) observing that the RING domain function of BRCA1 is not targeting of its substrates for degradation but instead promotion of protein localization and function at DNA damage sites, c) mechanistically elucidating how BRCA1 is recruited to and accumulates at DNA damage sites, and d) revealing that PALB2 mediates the interaction between BRCA1 and BRCA2 and functions in HR repair. Because PALB2, like BRCA1, is mutated in familial breast cancer patients, these data further confirmed the significance of HR repair in breast cancer suppression. This structural and functional knowledge of BRCA1 is essential to understanding the impact and significance of genetic mutations for breast cancer patients and clinical genetic counseling. These findings also have a major impact on the field of DNA damage signaling, as many other proteins involved in cell-cycle regulation, replication, and checkpoint control also contain BRCT domains. Moreover, the signaling pathway that recruits BRCA1 and the concept of degradation-independent functions of E3 ligase and ubiquitin as signaling moieties have foremost influence on studies of DNA damage response (DDR). We are now further investigating the biochemical activities of BRCA1, PALB2, and BRCA2 and attempting to mechanistically determine how these tumor suppressors act at different steps of HR repair. Moreover, we discovered several novel DDR-binding proteins. We will further investigate how these proteins participate in HR repair, genome maintenance, and tumor suppression in humans.
Besides our studies on BRCA1 and its roles in homologous recombination, we have also investigated many other DNA damage response and DNA repair pathways, which includes radiation-induced DNA double-strand break repair pathway, replication stress pathway and lesion bypass, fanconi anemia pathway and interstrand crosslink repair, Spartan/C1orf124/DVC1 and DNA-protein crosslink repair, and more recently the roles of DNA damage response in innate immunity.
Mass spectrometry-based proteomic analysis of tumor suppressive and oncogenic pathways
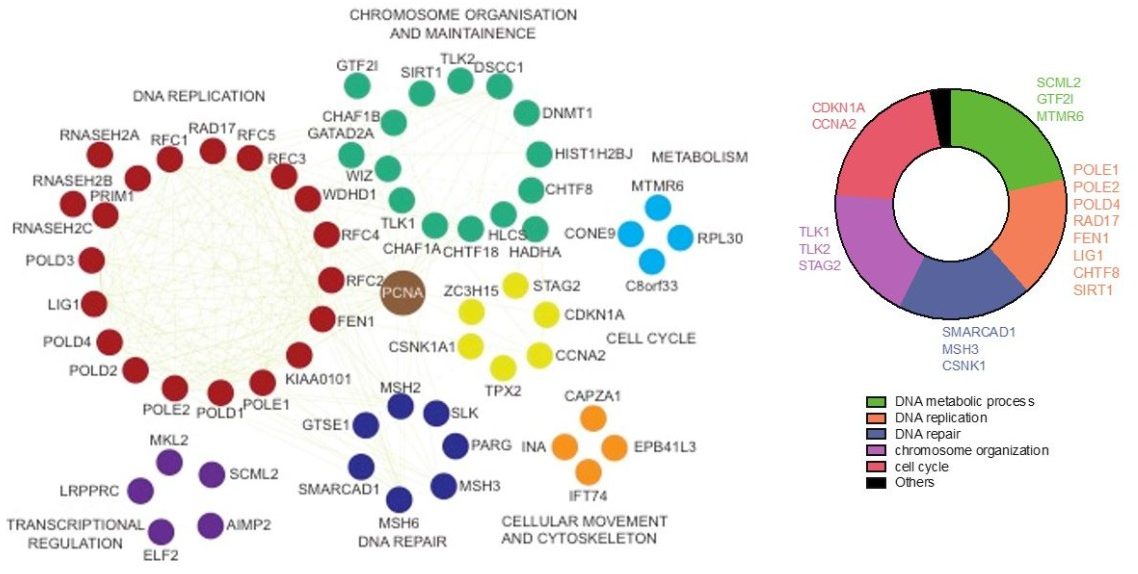
We employ several proteomic approaches for the studies of DNA damage repair pathways as well as tumor suppressive and oncogenic pathways. we have developed a novel tandem affinity purification protocol, which greatly facilitated our ability to isolate protein complexes from human cells. Over the years, we have used this new tandem affinity purification coupled with mass spectrometry (TAP-MS) approach for studies of DNA damage response and tumorigenesis. For example, we used this method to successfully identify a five-subunit complex, which consists of RAP80, CCDC98, BRCC45, BRCC36, and MERIT40, as BRCA1-associated proteins. In addition, we identified a previously uncharacterized nuclease, KIAA1018, now named FAN1 (Fanconi anemia-associated nuclease 1), that binds to mono-ubiquitinated FANCD2 via and participates in DNA repair in response to chemotherapeutic agents such as cisplatin and MMC. We also used this TAP-MS approach to conduct proteomic studies of several signaling pathways, for example, the Hippo/YAP pathway with 32 components. We completed a study of the CDK family (25 CDKs) in both MCF10A and HEK293T cells, which enabled us to identify common and specific CDK partners in these two cell lines. In addition, we conducted a proteomics analysis of 56 human transcription factors and demonstrated that transcription factors form distinct protein complexes on and off chromatin, which are important for the regulation and function of these transcription factors. More recently, we completed a study of mismatch repair protein interactomes and another study of protein phosphatase interactomes. All of these results testify to the success of our TAP-MS approach. Additionally, we have also adopted other technologies, such as BioID and APEX, which allow us to capture transient protein-protein interactions. Importantly, we developed a new bioinformatics algorithm and improved our ability to both identify and rank high-confidence interacting proteins (HCIPs), which enabled us to prioritize these HCIPs for further functional validation.
More recently, we are conducting quantitative mass spectrometry analyses using SILAC (stable isotope labeling with amino acids in cell culture), ITRAQ (Isotope Tags for Relative and Absolute Quantitation; post-labeling method, i.e. labeling peptides after proteins are digested by trypsin), and label-free quantitative mass spectrometry analysis. These quantitative mass spectrometry analyses can be used to address many biological questions. For example, we are studying the dynamics of DNA damage response by determining how DNA damage response network changes in response to treatment with a variety of DNA damaging and/or chemotherapeutic agents, including HU, radiation, topoisomerase poisons, cisplatin, and PARP inhibitors. In addition, we are using protein profiling and ubiquitinated proteome for the identification of key substrates of deubiquitinating enzymes and E3 ubiquitin ligases involved in DNA damage response, tumor suppressive and oncogenic pathways. Moreover, we are also studying phosphoproteome to investigate key cancer-related signaling pathways and response to targeted therapies. These studies will reveal new components and regulations involved in these cancer-related pathways and treatments.
CRISPR/Cas9-based genetic screens
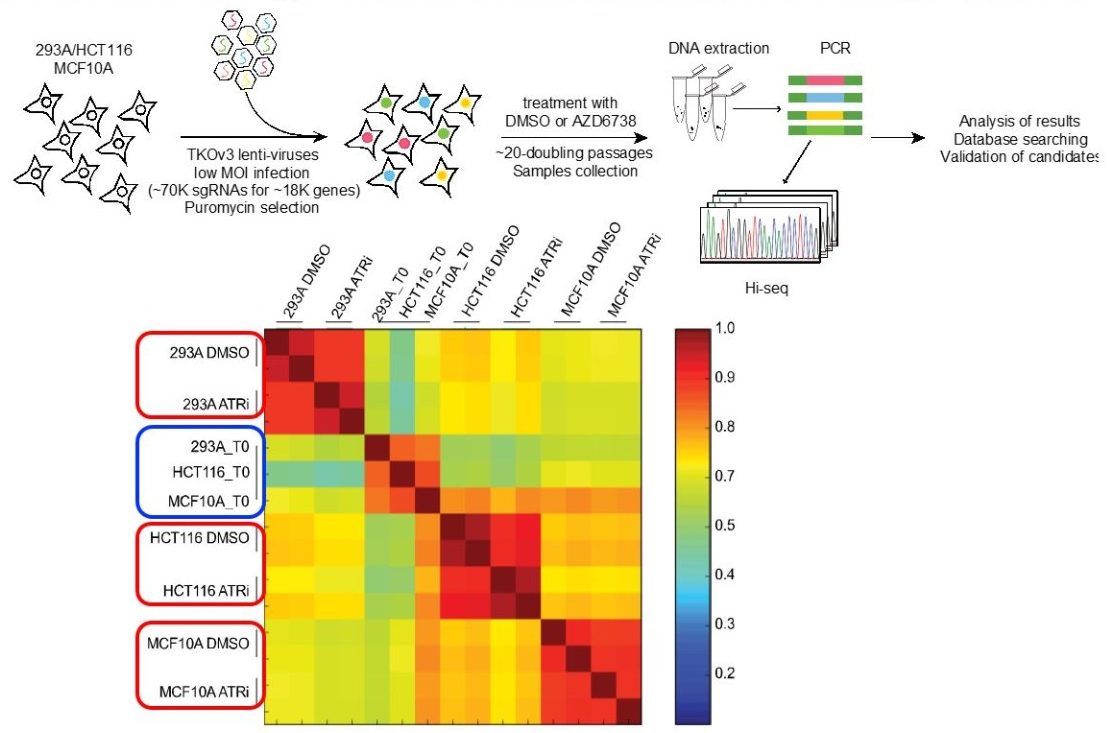
We are using CRISPR/Cas9-based gene editing approach to study functional redundancy in DNA damage signaling and other cancer-related signaling pathways. Moreover, we are performing genome-wide sgRNA screens for the discovery of gene-gene and gene-drug interactions.
We have started to generate panels of isogenic human cell lines, each with KO of a particular DNA repair gene, using a CRISPR/Cas9-based gene editing approach. This list includes genes involved in HR, NHEJ, FA, NER, MMR, BER, and TLS pathways. In addition, we have created a customized DDR sgRNA library which is being used to systematically determine genetic interactions between different repair genes and pathways. Moreover, we are performing unbiased whole genome sgRNA screens to discover genetic alternations that would sensitize these repair deficient cells (i.e. gene-gene interaction) or toward a given cancer therapy (i.e. gene-drug interaction). These screens are and will be conducted in two to three independent cell lines to reduce contribution of genetic variation in different cell lines. These genome-wide sgRNA screens are conducted in collaboration with Dr. Traver Hart, an experimental biologist and bioinformatics expert who is a faculty member in the Department of Biomedical and Computational Informatics, MD Anderson Cancer Center. Dr. Hart has designed, performed, and analyzed similar screens to identify fitness genes in multiple cell lines. In addition, he has further developed state-of-art algorithms for the identification of fitness genes in negative selection screens. Moreover, his lab also has extensive experience in identifying genes with variable knockout phenotypes across different backgrounds (e.g. different cell lines or isogenic knockouts) and different environmental conditions (e.g. +/- drug exposure). His experience and expertise are critical for the success of these whole-genome sgRNA screening studies. We anticipate that the close collaboration with Dr. Hart and the integrative analysis methods employed will redefine the state of the art for identifying novel targets for cancer treatment. We have successfully completed several of these screens and are geared up to perform additional gene-gene and gene-drug interaction screens. In addition, we will start in vivo screens using syngeneic mouse models.
Give Now
Research Areas
Find out about the four types of research taking place at UT MD Anderson.
