Cytogenetics and Cell Authentication Core
Asha S. Multani, Ph.D.
Director
- Research Resources
- Core Facilities and Services
- Cytogenetics and Cell Authentication Core
Overview
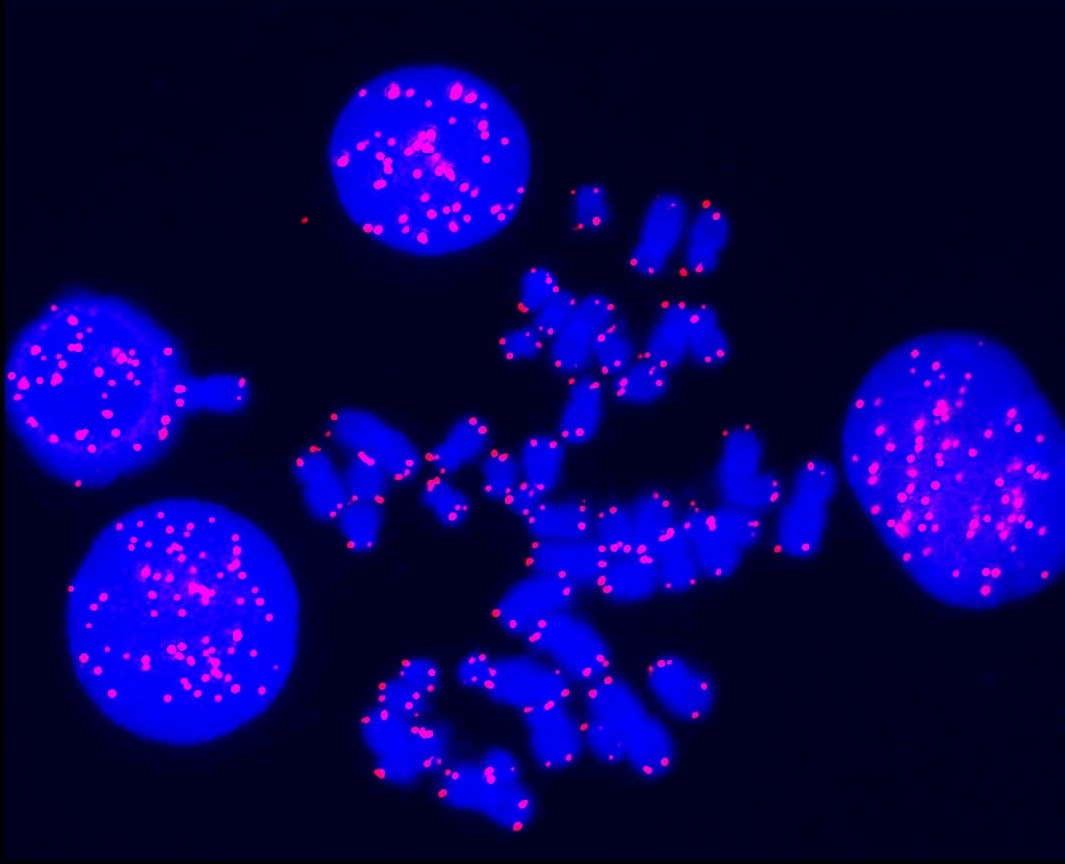
This facility offers conventional as well as molecular cytogenetic services including karyotyping, analysis of genomic instability, fluorescence in situ hybridization, telomere analysis, Spectral Karyotyping, species identification, inter-species and intra-species cell line contamination, STR fingerprinting service for cell line authentication, mycoplasma contamination testing and distribution of cell lines. Learn more about the core.
Getting Started
Please contact Dr. Multani to receive guidelines for sample preparation, technical assistance, to discuss the services that you require and to schedule an appointment.
Impact
The Cytogenetics and Cell Authentication Core (CCAC), since its inception in 2003, has contributed to numerous articles including several published in high impact journals, demonstrating the positive contribution of the CCAC shared resource to the advancement of science.
Contact
Asha S. Multani, Ph.D.
Associate Professor
713-563-1892
amultani@mdanderson.org
Facility Location
Mitchell Basic Science Research Building (BSRB)
S13.8414 and S13.8415 (lab)
S16.8316a (office)
Give Now
Research Areas
Find out about the four types of research taking place at UT MD Anderson.
